Ever since its discovery, the double-stranded DNA contained in the mitochondria of eukaryotes has fascinated researchers attributable to its bacterial endosymbiotic origin, important place in encoding subunits of the respiratory complexes, compact nature, and specific inheritance mechanisms.
In the last few years, high-throughput sequencing strategies have accelerated the sequencing of mitochondrial genomes (mitogenomes) and uncovered the great number of organizations, gene contents, and modes of replication and transcription found in dwelling eukaryotes.
Some early divergent lineages of unicellular eukaryotes retain certain synteny and gene content material materials resembling these seen in the genomes of alphaproteobacteria (the inferred closest dwelling group of mitochondria), whereas others tailor-made to anaerobic environments have drastically diminished and even misplaced the mitogenome. In the three main multicellular lineages of eukaryotes, mitogenomes have pursued quite a few evolutionary trajectories in which numerous sorts of molecules (spherical versus linear and single versus multipartite), gene constructions (with or with out self-splicing introns), gene contents, gene orders, genetic codes, and change RNA modifying mechanisms have been chosen.
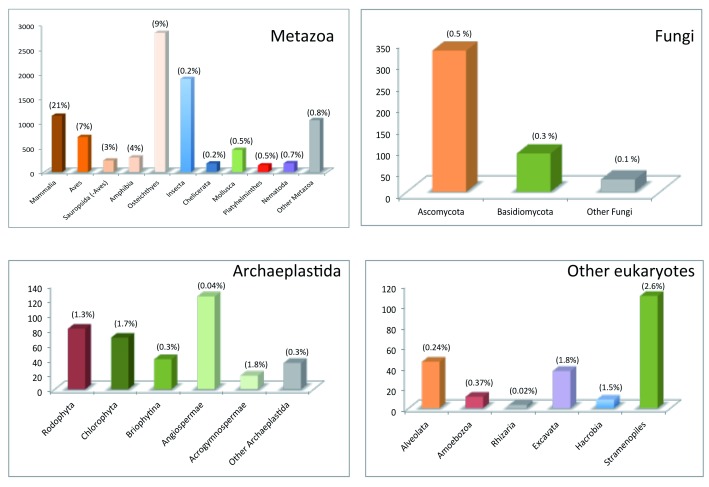
Whereas animals have developed a reasonably compact mitochondrial genome between 11 and 50 Kb in dimension with a extraordinarily conserved gene content material materials in bilaterians, crops exhibit large mitochondrial genomes of 66 Kb to 11.3 Mb with large intergenic repetitions liable to recombination, and fungal mitogenomes have intermediate sizes of 12 to 236 Kb.
Genome diversification in globally distributed novel marine Proteobacteria is linked to environmental adaptation.
Proteobacteria signify one of many essential quite a few and ample groups of microbes on Earth. In productive marine environments like deep-sea hydrothermal packages, Proteobacteria are implicated in autotrophy coupled to sulfur, methane, and hydrogen oxidation, sulfate low cost, and denitrification.
Beyond chemoautotrophy, little is believed in regards to the ecological significance of poorly studied Proteobacteria lineages which will be globally distributed and energetic in hydrothermal packages.
Here we apply multi-omics to characterize 51 metagenome-assembled genomes from three hydrothermal vent plumes in the Pacific and Atlantic Oceans which will be affiliated with 9 Proteobacteria lineages.
Metabolic analyses revealed these organisms to comprise a numerous purposeful repertoire along with chemolithotrophic functionality to profit from sulfur and C1 compounds, and chemoorganotrophic functionality to profit from environment-derived fatty acids, aromatics, carbohydrates, and peptides. Comparative genomics with marine and terrestrial microbiomes signifies that lineage-associated purposeful traits would possibly make clear space of curiosity specificity.
Our outcomes clarify the ecological options and metabolic strategies of novel Proteobacteria in hydrothermal packages and previous, and highlight the connection between genome diversification and environmental adaptation.
